Background
You are what you eat, or so many people have been told. Researchers today are discovering that this age-old saying may be even more true that previously thought. “The Human Microbiome Project (HMP) was a United States National Institutes of Health (NIH) research initiative to improve understanding of the microbial flora involved in human health and disease’(7). Actually, the human microbiome project is a continuing effort of scientists from across the globe working together to better understand how all of the microbiological members of the immediate human environment (the microbiome) interact with our daily lives. The human microbiome is vast network of trillions of microorganisms that live in and on the human body including bacteria, fungi, parasites, and viruses(9). According to the National Institute of Health’s website, the human body is made up of 1-3% of microorganisms by mass(1).
That number has since been debated amongst scientists on being much larger and thus the effects of our bodies cohabitants may be more significant than we previously anticipated. This really does bring new validity to referring to one’s own body as “our body’. Some microorganisms may produce compounds that our body puts to use, some may only interact with other microbiota, and others may simply live in our bodies providing little to no interaction whatsoever. Finding out which microorganisms affect our bodies in favorable and unfavorable ways is something researchers are constantly making progress on. The process of qualifying the functions of all these different members of our microbiome communities can be complex and challenging.
The modern method of detecting the presence of microorganisms in the body, commonly referred to as microbiota, is through a technique called 16S (16S is a gene) metagenomic sequencing. This is a fancy way of saying geneticists look for specific parts of RNA that are exclusive to organisms so we can identify them individually. RNA (Ribonucleic acid) is a polymeric molecule essential in various biological roles such as the coding, decoding, regulation and expression of genes(10). Genes are small sections of DNA within the genome that code for proteins. They contain the instructions for our individual characteristics — like eye and hair color(11). Scientists will take samples from subjects and analyze the RNA in those samples through 16s sequencing and then compile a list of which microorganisms are present and in what amount or abundance they occur. By knowing this information researchers can then compare the results to a broad range of human health factors including hypertension, diabetes, insomnia, and many more in an attempt to find potential links between our microbiota and our health.
A study recently conducted at the University of Wisconsin Madison (UMW) called “Gut microbiome populations are associated with structure-specific changes in white matter architecture’ shows how biostaticians (scientists who apply statistical methods to analyze topics in biological) and radiologists have teamed up to better understand the link between the gut microbiota and neurological tissue structure of rats(2). The brain’s white matter is given its name because of the myelin which gives it its color. Myelin protects nerve fibers from injury and also improve the speed and transmission of electrical nerve signals(3). The changing of gut microbiota has been shown to modify behavioral function in animals. In this study scientists aim to better understand the connection between changes in behavior, changes in gut microbiota, and changes in brain structure. This may allow us to gather new insights into the function of our own gut-brain axis. The gut-brain axis refers to the connection between the microorganisms in the human digestive system and the central nervous system(4).
Evidence
Young male rats given different diet treatments ( Standard diet, high fat diet, high fiber diet, and high protein low carb diet) were shown to have measurably different brain architecture. In other words what the rats ate had a specific effect on various areas of their brain’s development. As the following figure 1 displays, modern imaging techniques can illuminate the neural (brain tissue) differences we want to study in relation to different diets and different gut microbiomes.
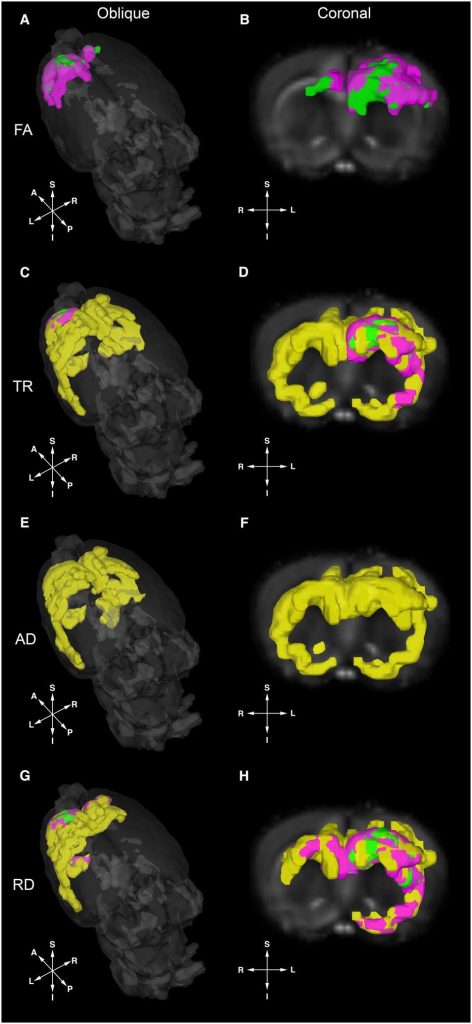
Fecal samples of the rats were taken both initially and three weeks after the dietary treatments were applied to see how different diets affected the make-up of their gut microbiomes. As would be expected, if microbiomes were indeed influenced by dietary choices, there were significant differences in the populations of bacteria for rats with different diets. Not only were there varying amounts of bacteria, but there were unique gut communities specific to each diet. Based on these findings the researchers were able to come up with a way to predict the diet treatment category of a rat based on their gut microbiome. The next step was finding out whether or not these predictions, based on gut microbiomes, could be predictive of the brain architectural changes observed. Unique structural regions of interest in the brains of the rats were compared and the apparent differences were able to be used to discriminate between the diet treatments with 95% accuracy.
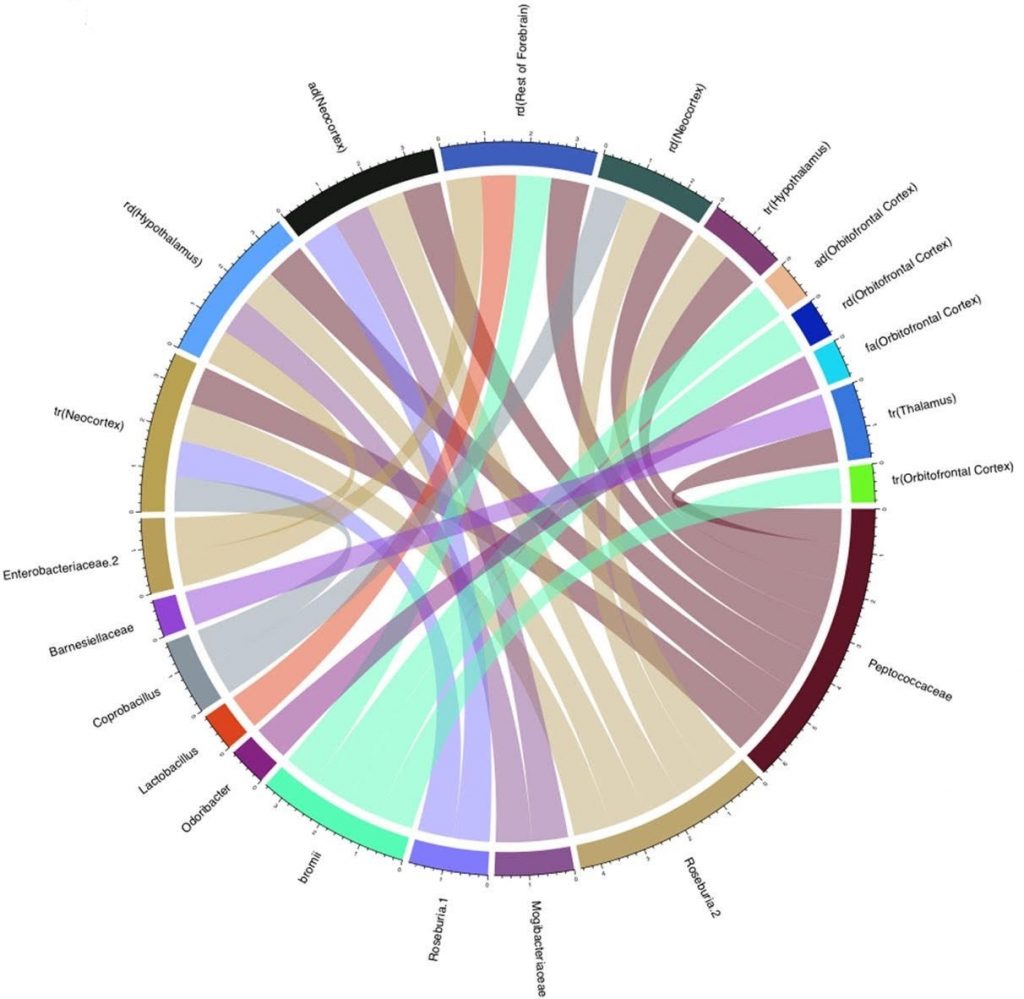
The next step in this study was to make comparisons between the gut microbiomes of the rats, diets, and the brain structure regions of interest to create a predictive mechanism of finding specific links between bacterial communities and brain architectural differences. The following Figure 2 labels the names of bacteria which were shown to uniquely affect areas of the rats brains. Knowing the mechanisms of how these gut populations develop can help illuminate how neural changes may come about as a result of dietary choices.
Questions
You may be wondering if comparing the brain imaging of humans and their respective gut microbiomes will show us specific influences that our guts have on our minds. People who choose to eat different foods may cause changes in their gut microbiomes which may then cause changes in their brains. These neural changes can then have both long term and short term effects on their behavior. If we can take a closer look and find a connection between what people eat, their microbiome, and their behaviors we can begin to use this knowledge to improve health and hopefully prevent the development of undesirable influences on our brain from our food. If our gut really does have control over our mind, then the phrase “you are what you eat’ is true in more than one way.
Conclusion
This kind of research has led scientists to believe that although the biological connection of the gut microbiome to the “central nervous system has several effects’ there lies specific changes in brain structure associated with the composition of gut microbiota(2). Our gut community produces a wide variety of compounds including neurotransmitters which can directly affect the function of our brains(8). Finding specific links between gut microbiota members and structural changes in the brain can help us understand how the choices we make can play a role in helping or hindering our neural health by interacting with our gut microbiota composition. This can also shed light on potential mechanisms of neurodevelopmental disorders.
Studies like these are a never-ending avenue for discovering new links in the complex mystery of what makes us human, and as it turns out, a lot of our behavior may be due to bacteria! So the next time you make a decision, whether you go with your brain or your gut, it may be your gut’s choice anyhow!
Further Reading
The following supplementary articles are directly related to the research performed in this study and are aimed at better understanding the link between the gut microbiome and how neural development & function can be closely related.
- The Microbiota-Gut-Brain Axis (4)
- The Gut—Brain Axis and the Microbiome: Mechanisms and Clinical Implications (5)
- Crosstalk Between the Gut Microbiota and the Brain: An Update on Neuroimaging Findings (6)
References # (1-11)
- NIH Human Microbiome Project defines normal bacterial makeup of the body. (2015, August 31). Retrieved from https://www.nih.gov/news-events/news-releases/nih-human-microbiome-project-defines-normal-bacterial-makeup-body.
- Ong, I. M., Gonzalez, J. G., McIlwain, S. J., Sawin, E. A., Schoen, A. J., Adluru, N., … Yu, J.-P. J. (2018, January 10). Gut microbiome populations are associated with structure-specific changes in white matter architecture. Retrieved from https://doi.org/10.1038/s41398-017-0022-5
- White matter of the brain: MedlinePlus Medical Encyclopedia. (n.d.). Retrieved from https://medlineplus.gov/ency/article/002344.htm.
- Cryan, J. F., Oriordan, K. J., Cowan, C. S. M., Sandhu, K. V., Bastiaanssen, T. F. S., Boehme, M., … Dinan, T. G. (2019). The Microbiota-Gut-Brain Axis. Physiological Reviews, 99(4), 1877—2013. doi: 10.1152/physrev.00018.2018
- R.Martin, C. (2018, October 4). The Gut—Brain Axis and the Microbiome: Mechanisms and Clinical Implications. Retrieved from https://doi.org/10.1016/j.cgh.2018.10.002
- Liu, Ping, Peng, Guoping, Zhang, Ning, … Benyan. (2019, July 30). Crosstalk Between the Gut Microbiota and the Brain: An Update on Neuroimaging Findings. Retrieved from https://doi.org/10.3389/fneur.2019.00883
- Human Microbiome Project. (2019, September 23). Retrieved from https://en.wikipedia.org/wiki/Human_Microbiome_Project.
- PhilipStrandwitz. (2018, June 11). Neurotransmitter modulation by the gut microbiota. Retrieved from https://doi.org/10.1016/j.brainres.2018.03.015
- The Microbiome. (2019, September 4). Retrieved from https://www.hsph.harvard.edu/nutritionsource/microbiome/.
- RNA. (2019, October 26). Retrieved from https://en.wikipedia.org/wiki/RNA.
- What is a gene? (2016, October 6). Retrieved from https://www.yourgenome.org/facts/what-is-a-gene.
